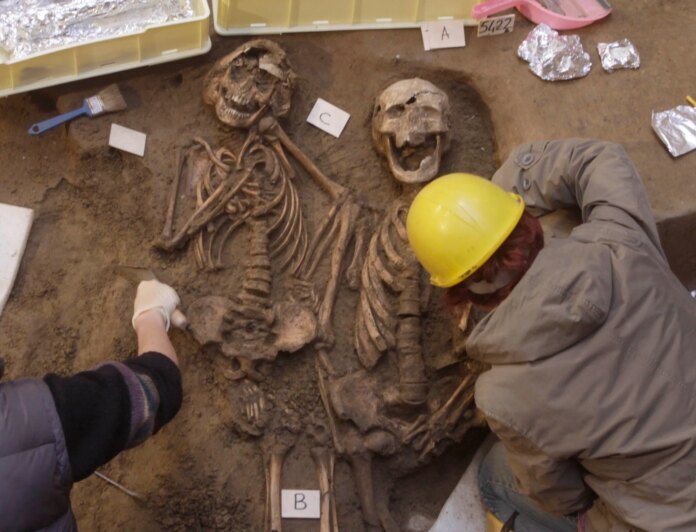
Some 4,000 years ago, as ancient civilizations such as the Minoans in Crete and the Neo-Sumerian Empire in Mesopotamia were shaping cultures in Europe and the Middle East, human biology itself was evolving apace.
A groundbreaking study of ancient DNA, published last week in the prestigious journal Nature, has documented that the frequency of two DNA variants linked to celiac disease increased dramatically across populations in those regions.
Hundreds more such DNA variants were also uncovered in the study, which sheds unprecedented light on how, in the past 10,000 years, human evolution, and specifically the biology of the human body, have been shaped by natural selection. (Natural selection occurs when a version of a gene linked to a specific trait, such as a disease like celiac, but also qualities like red hair or a blood type, proves advantageous enough for survival and reproduction).
Led by scientists from Harvard University, the researchers analyzed and compared the genomes, or complete DNA sequences, of some 22,000 individuals. The data included 10,000 ancient genomes never studied before, 6,000 previously published genomes, and 6,000 modern individuals.
Previously, only 21 genetic shifts due to natural selection — as opposed to migrations, community mergers, or similar factors — were known.
“It’s so powerful to be able to watch evolution happening in action, not just to study the scars that evolution leaves on modern patterns of variation,” the study’s senior author, Harvard geneticist David Reich, told The Times of Israel in a video interview.
‘Industrializing the production of ancient DNA
Reich, who heads a lab focused on ancient DNA, biology, and disease, explained that extracting ancient DNA has only been possible for the past few years, and it took time to accumulate enough data to conduct a study on human evolution.
“The technology for getting ancient DNA out of human remains only became available beginning in 2010,” he said. “I think it’s not an understatement to say that it has had a transformative impact on our understanding of the past.”
The scientist has authored many studies on the human history of specific groups, archaeological sites, and regions, including Jews and populations in the land of Israel.
The Nature paper, which analyzed genomes from countries spanning from Iceland to Spain, Russia, Iran, and Israel, took a different approach, investigating human biology.
“Many of the people who started this field were biologists, and really thought that the most exciting thing one could do with ancient DNA would be to try to understand how our biology might have changed over time, ” he said. “However, in order to study human history, you only need one or a few samples per population, but to understand evolution, you need many samples. The data just did not exist.”

To make this type of study possible, the Reich Lab and other laboratories worldwide focused on “industrializing the production of ancient DNA” for the last decade, according to the scientist.
“We extracted the DNA from individuals with robots, cleaned it up with robots, and we turned it into a form that can be sequenced with robots,” he said. “We also used all sorts of computational methods to analyze the data in uniform ways that produce high-quality, good data.”
The Reich Lab collaborated with some 270 archaeologists who provided samples with the basic data needed for the analysis, with the understanding that research touching on the study’s human-history angle would be shared later.
“The archaeologists are not particularly interested in the biological change question, and they saw that the data could be useful and could be released in a way that did not detract from the story about the archaeology,” Reich said.
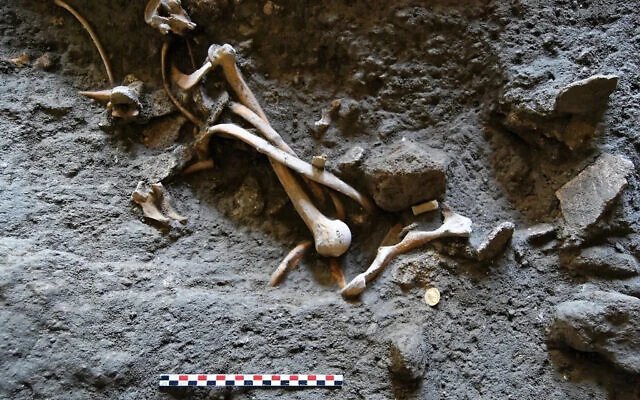

As a result, according to the geneticist, the new study has doubled the amount of data in the entire field of ancient DNA — and tripled what was available for West Eurasia.
Asked how the ancient samples were dated, Reich said they only worked with samples they felt had reliable, relatively precise dating, either based on radiocarbon dating, clear placement in a timeline based on the layer of soil in which it was found, or a combination of both.
“We excluded samples that had less clear chronology,” he said.
Going beyond population movements
Another challenge the paper solves, thanks to the intuition of the study’s lead author, Reich Lab senior staff scientist Ali Akbari, is documenting only shifts caused by natural selection alone.
“History has been so complicated everywhere in the world, including in Europe and the Middle East, with lots of population movements and migrations and mixtures,” Reich said. “Many of the changes are just due to the fact that new people arrived in a place, or they resulted from a mixture of two groups. We needed a strategy to detect the needle in the haystack.”
The statistical model the researchers used compared each genome to the other 22,000 and measured how closely related they were.
“Most human genomes are extremely similar to each other, so if you line them up with each other, 99.9 percent of the DNA letters will be the same,” Reich explained. “Out of 3 billion letters, which is the length of a genome, there might be 3 million differences, which sounds like a lot, but really represents only one out of 1,000 of the whole.”

The system analyzed approximately 10 million letters that differed across the human genomes studied and examined whether considering time alone helped predict changes.
“The previous literature had found only about 20 positions [in the genome] that had changed significantly over time,” said Reich. “Now this has increased to almost 500 such positions where we find clear evidence of adaptation, and probably several thousands more, if you lower your threshold for certainty.”
The stark increase in the frequency of the two DNA variants associated with celiac disease risk was among them.
“You might have thought that these variants would have become less common in the last few thousand years because, as people started eating wheat, [they could have] become less susceptible to these things, but they became more susceptible,” Reich said. “Why? We don’t know. Probably because [these variants] also protected against some other disease.”

The scientists also documented a shift connected with the risk of contracting tuberculosis.
“There are several genetic variations [whose frequency] kind of flipped,” Reich said. “One of these variations is the most important risk factor for serious cases of tuberculosis. We see very strong natural selection for the risk factor to increase in frequency between 6,000 years ago and 2,000 years ago. Afterward, there was a very strong selection against it.”
“Speculatively, what happened was that tuberculosis became widespread, and carrying the variant became very bad, but before that, it may have protected against something else, another infectious disease, so it was good to carry it,” he added.

The study documented the evolution of variants associated with a range of conditions, including multiple sclerosis, hemochromatosis, bipolar disease, and Crohn’s, as well as with human traits such as red hair, light skin tone, and different blood types.
The authors also published an open database where any fellow researcher can look up any location in the human genome and see which DNA variants are found there.
Asked whether the study could offer new insights into ancient Jewish DNA or specific genetic variants associated with diseases common in the Jewish community, Reich said there were not many samples from Jewish individuals included in the study.
He said ancient DNA from Jewish individuals is generally easy to identify, as Jews historically tended not to mix with surrounding populations. However, because Jews are typically buried in distinct cemeteries, the likelihood of identifying Jewish DNA in a non-Jewish burial context is low.

Similarly, the scientists did not examine how specific regions or populations within West Eurasia compared with one another, including specifically investigating ancient DNA from the land of Israel.
However, this will likely be an avenue for future research, as Reich emphasized the current study is “only the beginning,” highlighting the potential for its data and results to be used in further studies across the medical and biological fields, as well as in history and archaeology.
“We don’t have any systematic observations yet that distinguish Near Eastern populations from European populations, but that’s an important and interesting direction,” he said.
Further expanding the pool of genomes analyzed could also yield important results.
“We worked with an amazing data set that can be reused for all sorts of purposes, but it’s very clear that if you further increase the sample size, you’ll get more discoveries,” he said.
In the future, the scientist also hopes to conduct a similar study in other regions of the world.
“What results could we get if we conducted a study like this in East Asia, or among Native Americans? Would they be similar or different?” he noted.
Reich explained that similar research could also be conducted on animal DNA to better understand their evolution.
“These are the things that we’re interested in doing and could be interesting to do next,” he said.
Source:
www.timesofisrael.com


